 Note that uniq won't detect repeated lines in the input file if they are not adjacent, so it may be necessary to sort the file. How can I do this with bash? Possibly listing the number of occurrences next to an ip, such as: 5.135.134.16 count: 5 I want to find the number of occurrences of each unique IP address. The output of awk can then be piped into "LC_ALL=C sort -n | uniq -c" to get the read size distribution.I have a log file sorted by IP addresses, The statement "if (NR%4 = 2) print length($0)" means "output the size of the second line in every sequence record". The expression (NR%4) gives you the remainder of NR divided by 4. The "awk" function has a variable "NR" that records the line number for each row. To reduce the number of results that are displayed, use the -m (max count) option. The line number for each matching line is displayed at the start of the line. In a fastq file, each sequence record has 4 lines and the second line of the record is the actual DNA sequence. You can make grep display the line number for each matching line by using the -n (line number) option.  Äœalculate the read length distribution.In this exercise, you will use a pretty old tool called fastx_clipper, and write the output into a new fastq file "clean.fastq". The hash keys are the elements of greps input list the hash values are running counts of how many times an element has passed through greps BLOCK. preggrep preglasterrormsg preglasterror pregmatchall preg. To get the number of occurrences of each unique value, use uniqs -c option: sort mylist. In that case, you might want to use reads from the middle of the file by using "head -n xxxxx | tail -n xxxxx" command) count) echo count //3 > The above example will output: xpto 3. That will give you the number of unique values. 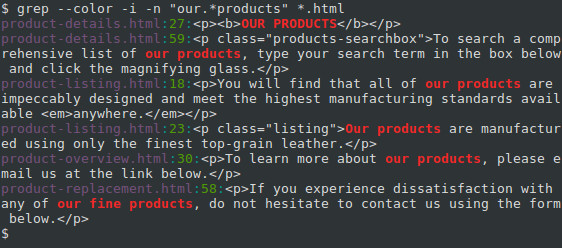 Sometimes the first 10,000 reads are all low quality reads, then this estimation would not be accurate. You can use the first 10,000 sequencing reads to estimate. Grep or awk a unique and specific word across many fields Hi there, I have data with similar structure as this: CHR START-SNP END-SNP REF ALT PATIENT1 PATIENT2 PATIENT3 PATIENT4 chr1 69511 69511 A G homo hetero homo hetero chr2 69513 69513 T C. In case you are using git, the command git grep -h sort -unique will give unique occurrences of grep matches.The one marked as duplicate is different because it it is not about grep.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
